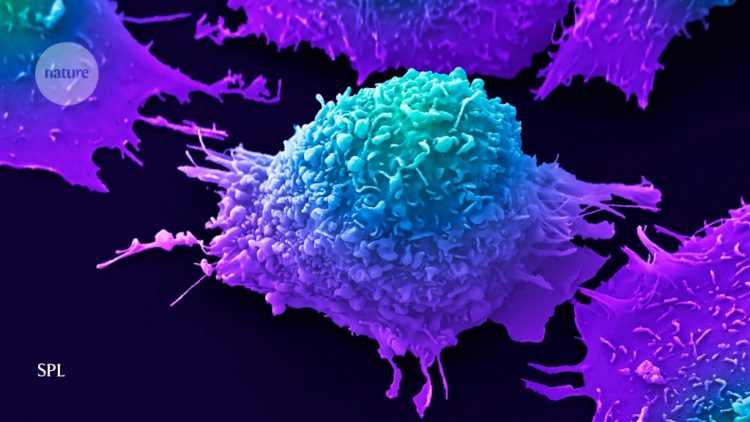
By sifting through hundreds of millions of mutations lurking in the genomes of more than 12,000 tumours, researchers have identified patterns of DNA changes that could offer clues to the genetic and environmental causes of cancer.
The study, published in Science on 21 April1, is the largest of its kind. It adds dozens of entries to the growing catalogue of ‘mutational signatures’ that accompany cancer, and could, in some cases, help clinicians to select the best treatments for individuals.
Size matters for these analyses, says Núria López-Bigas, a computational cancer biologist at the Institute for Research in Biomedicine in Barcelona, Spain. The new work has revealed rare mutational patterns that could not have been picked out from smaller data sets. “When you have this number of whole genomes, you have more power and can make a more complete set of mutational signatures,” she says. “It is still early days, but it has a lot of potential in diagnosis and to understand how these tumours have been created.”
Mutational footprints
An individual cancer cell can contain hundreds of thousands of mutations, sometimes more than one million, but only a handful of these will contribute directly to the development of a tumour. For years, researchers have been trawling through genomic data in search of these cancer drivers, in the hope that they could point to new therapies.
The many remaining ‘bystander’ mutations can also be informative. Some cancer-causing agents create characteristic patterns of DNA changes. Ultraviolet light, for example, can cause a DNA base, or ‘letter’, called cytosine to be replaced by another called thymine at certain sites in the genome. Such changes are often found in melanomas.
These patterns of mutations can be likened to footprints on a sandy beach, says Serena Nik-Zainal, a computational biologist at the University of Cambridge, UK, and a co-author on the Science study. “The footprints may look random, but they are not — they are occurring for a very particular reason,” she says. “You would be able to distinguish a human from an animal, a dog from a bird, even an adult from a child and whether they were walking or running.”
The largest previously reported study of mutational signatures was published in 2020 and analysed about 5,000 whole-genome sequences from tumour samples collected in an international effort2.
In the new study, the team analysed more than 12,000 cancer genomes collected by the UK National Health Service as part of the Genomics England 100,000 Genomes Project. The researchers then used previously published data sets to verify their findings. This involved developing new analytical tools and an algorithm capable of handling hundreds of thousands of mutations, says Andrea Degasperi, a computational biologist at the University of Cambridge and a co- author on the study.
The work — which included samples from 19 tumour types — yielded dozens of previously unknown mutational footprints, some of which could be traced back to defects in specific cellular methods for repairing DNA.
Researchers have probably now found all of the most common mutational signatures, says Dávid Szüts, a cancer biologist at the Research Centre for Natural Sciences in Budapest. “It seems unlikely that the major processes are missed at this point,” he says. But the hunt for rare signatures that occur in less than 1% of tumours in a given organ will probably continue as cancer-genome projects flourish worldwide.
In addition to searching for further mutational signatures, Degasperi hopes to be able to track down the origins of the more mysterious ones that have not yet been linked to a cancer-causing event. He also wants to investigate other kinds of genetic change: the current study focuses on changes to between one and three DNA letters, but DNA sequences can also be deleted, inserted or rearranged in larger chunks.
The hope is that these studies will eventually lead to cancer treatments that are tailored to individual people, he says. “When you understand the mechanism, you might understand a possible correlation with a drug.”